While many of the posts on this blog are written with an audience of medicinal and computational chemists in mind, this one is going to be a little bit of a departure. We’ll be exploring a fun math problem we ran into while pursuing some assay design work, and going into some details about algorithms that may be of interest to the computer science crowd. For readers who are newer to drug discovery, we hope this gives you a taste of some of the interesting constraints and challenges at the intersection of machine learning and wet lab science. But whatever your background, we’ll try to keep things accessible; you don’t need to know abstract algebra to follow along. And if you do know a bit about group theory, we hope you’ll find the application interesting!
The setup for this problem involves pooling small molecule compounds to increase throughput in experimental assays. Typically compounds are prepared for measurement by dissolving them in a solvent and placing them in a series of wells in what’s called a plate. Pooling, in this context, means mixing multiple different compounds into a single well, and running the measurement on all the compounds at once. As a specific example, we might want to measure how rapidly a molecule is metabolized by liver enzymes. With pooling, we expose a number of compounds to liver enzymes in the same well at the same time, and measure individual results for each compound at the end. At Inductive this is something we've been exploring as a way to acquire data more efficiently, and no machine learning practitioner has ever said no to more data!
When pooling compounds, though, you always run the risk that the presence of other compounds in the same well will somehow affect the results ("drug-drug interactions"). In the case of metabolic stability, one way this can happen is that some drugs will inhibit the metabolizing enzymes–which will affect how rapidly other compounds in the pool will be metabolized. To mitigate this risk, we wanted to design a pooling scheme such that each compound is measured in multiple different pools, so that we can cross-check the results for potential drug-drug interactions. When the results for a compound from all pools agree, we can be confident that the measurements for that compound are robust. In order to identify potential interactions, though, we need the compounds involved to be measured in other pools where they won't interact. The problem that arises, then, is how to design those pools such that we minimize the number of compounds that always appear together in the same pool. We can formulate this problem precisely with some math, and the rest of this post will describe how we can do that and some results that follow.
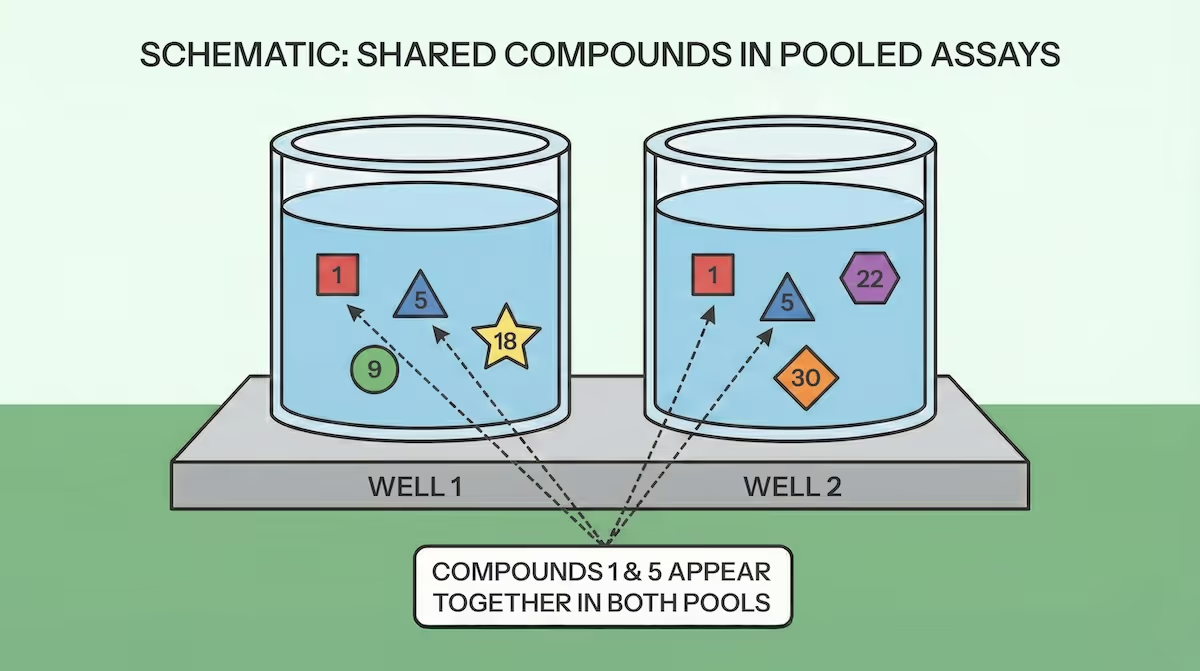
Suppose we have a plate with $W$ wells (numbered $0$ to $W-1$), and we want to place $C$ compounds (numbered $0$ to $C-1$) into those wells. We will measure $R$ replicates of each compound, so each compound will be present in $R$ different wells. This means each well will contain $P$ pooled compounds, and we have $CR=PW$. (Note that we don't need to constrain $P$ to be an integer, we might have some wells with extra compounds to cover the remainder.) We can then define a matrix $M_{w,c}$ that is $1$ if compound $c$ is present in well $w$ and $0$ otherwise. Our goal is to design the placement of compounds into wells such that the number of compounds that always appear together in the same wells is minimized, or:
$$\min \sum_{c=0}^{C-1} \sum_{c'=0}^{c-1} \prod_{w}[M_{w,c} = M_{w,c'}]$$
Before getting to the optimal solution, let's first consider a straightforward approach. For constrained optimization problems like this, a random algorithm is generally a useful baseline. We can write some simple python code to test this out:
import random
def random_fill(num_wells, num_compounds, num_replicates):
plate = [[] for _ in range(num_wells)]
for cpd in range(num_compounds):
used_wells = []
for _ in range(num_replicates):
valid_wells = [i for i in range(num_wells) if i not in used_wells]
min_fill = min(len(plate[i]) for i in valid_wells)
chosen_well = random.choice(
[i for i in valid_wells if len(plate[i]) == min_fill]
)
plate[chosen_well].append(cpd)
used_wells.append(chosen_well)
return plateThis will fill our plate compound by compound, trying to keep the wells as evenly filled as possible. We can test this out numerically for a few different parameter settings to see how well it performs. Below we've plotted some results for $W=24$, $R=2$ and $R=3$ replicates, varying the number of compounds from 0 to 500. We're plotting the average number of compound pairs that always appear together in the same wells over 100 random trials:
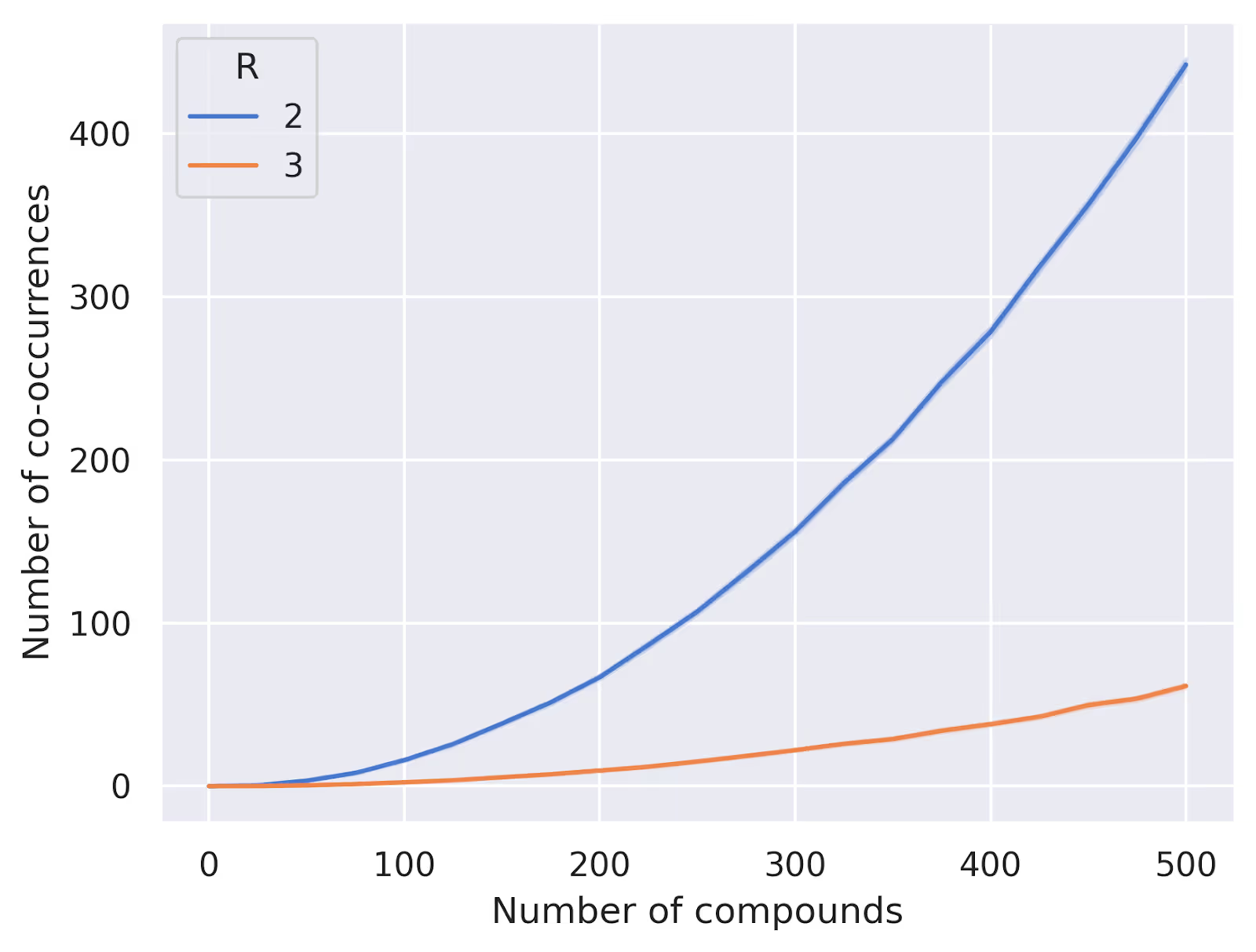
How far is it from optimal? Well, let's think a bit more about the constraints of our problem. When we place a compound into our plate, we are selecting $R$ wells out of the total $W$ wells without replacement. This corresponds to the mathematical concept of combinations, and there are thus a total of $\binom{W}{R} = \frac{W!}{(W-R)!\,R!}$ possible unique patterns of well placement for each compound. If we have $C$ compounds to place, we can only guarantee a unique pattern for each compound if $C \le \binom{W}{R}$. From the plot above, we can see that our random algorithm starts to give us duplicates long before the number of compounds reaches this limit of $276$ for $W=24$ and $R=2$.
This insight about enumerating combinations leads us to a natural solution: we can iterate through the combinations of $R$ wells out of $W$ (multiple times if needed), and assign each compound to a different combination of wells. This solution is optimal—if the number of compounds happens to be an integer multiple of $\binom{W}{R}$. Practically, however, this is unlikely to be the case: if we revisit the case of $W=24$ and $R=2$, it's not very convenient to require multiples of 276 compounds. And if we try to use partial iterations through the list of combinations, we run into a problem:
>>> from itertools import combinations
>>> list(combinations(range(24),2))[:10]
[(0, 1), (0, 2), (0, 3), (0, 4), (0, 5), (0, 6), (0, 7), (0, 8), (0, 9), (0, 10)]Note that all the combinations at the beginning include 0. This list of combinations may fill the wells very unevenly if we stop partway through. The plot below shows the difference between the number of compounds in the most and least filled well using this strategy:
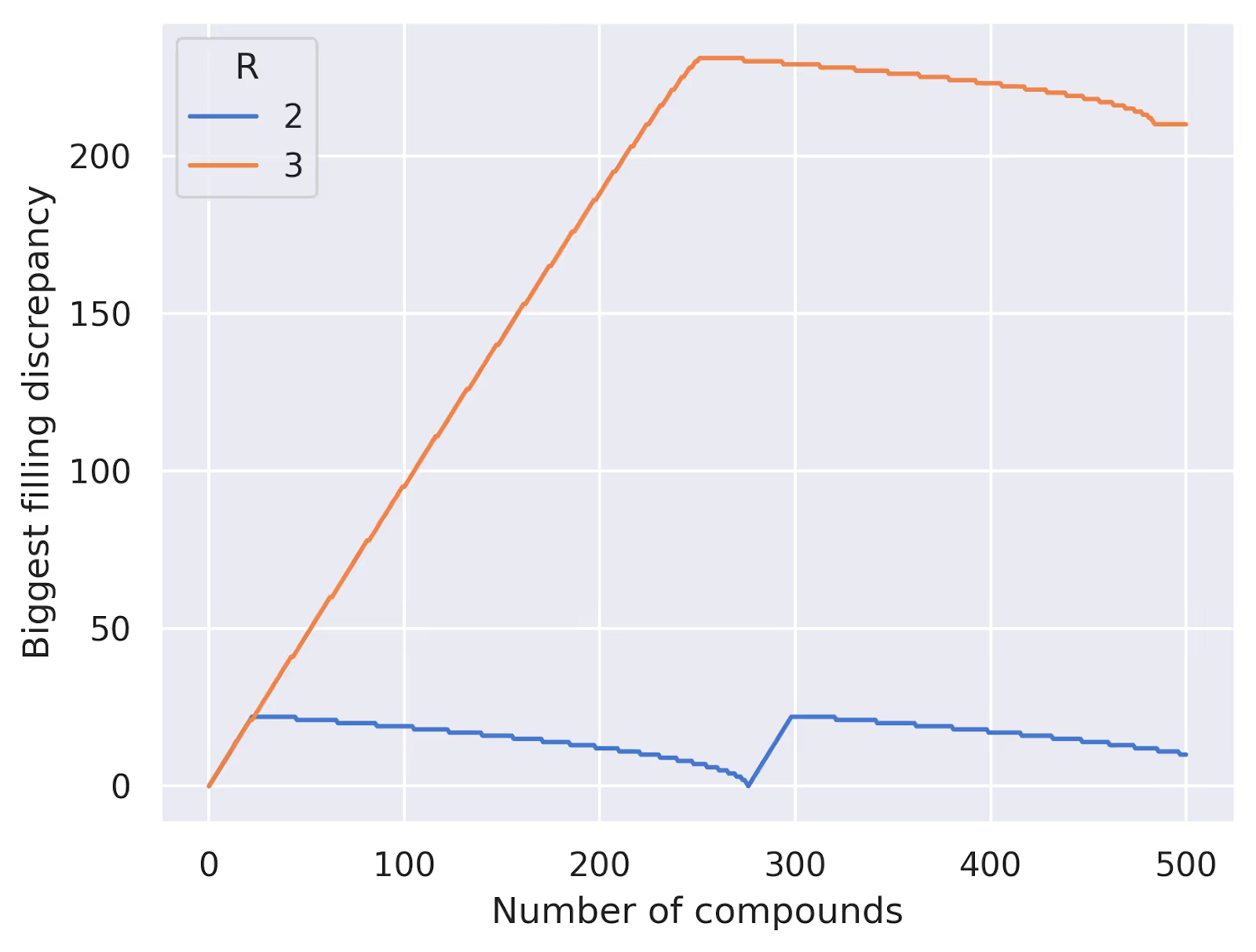
What we want, then, is to somehow order the set of unique combinations such that we'll also fill the pools evenly as we go. I'd encourage you to take a moment to think about how you might do this before reading on!
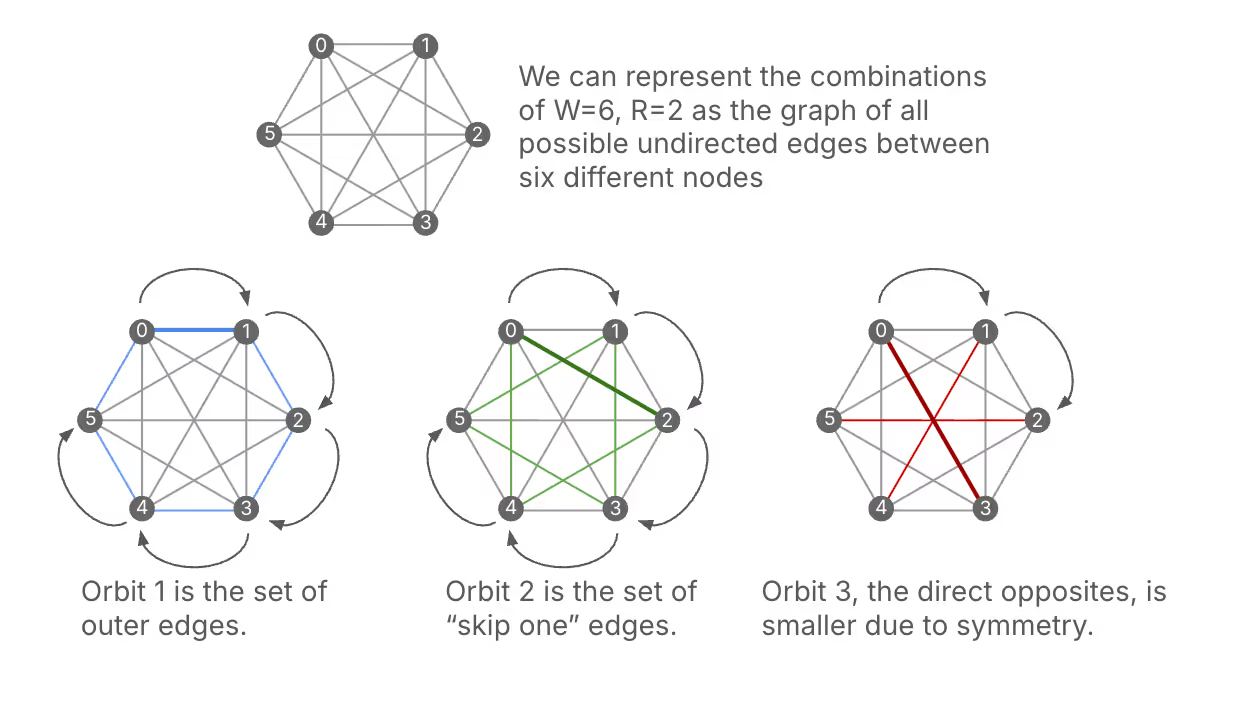
Let's first walk through a simple case, where $W=6$ and $R=2$, to get a feel for how we can do this. We'll take the following approach: first, start with the combination {0,1}. Then, we'll add 1 to each element, modulo 6, to get the next combination {1,2}. We'll keep doing this until we return to {0,1}. This gives us the following sequence of combinations: {0,1}, {1,2}, {2,3}, {3,4}, {4,5}, {5,0} Notice that in this sequence, each well appears exactly twice, so this subset of combinations will fill the wells evenly. Great! Now, we can start with a new combination that hasn't been used yet, say {0,2}, and repeat the same process: {0,2}, {1,3}, {2,4}, {3,5}, {4,0}, {5,1} Again, each well appears exactly twice in this sequence.
At this point, you might be asking if this approach is guaranteed to work. In particular, how do we know that we will exhaustively visit all combinations without getting any repetitions? Well, it turns out that this approach can be described with the language of group theory, which gives us some powerful guarantees on our results. If we think of the possible combinations of wells as a set, we can say that we're applying a group action of the cyclic group of order $W$ on the set of combinations. Each subset of combinations generated in this way is called an orbit under this group action. One result we can draw from group theory is the orbit decomposition theorem, which tells us that the set of all combinations can be decomposed into disjoint subsets via these orbits. This means that by applying our method, we can systematically generate all combinations while ensuring even filling of the wells.
Further, a result called the orbit stabilizer theorem tells us more about the sizes of the orbits that we generate. Specifically, the size of each orbit is equal to the size of the group (which is $W$) divided by the size of something called the stabilizer subgroup of the starting combination. In practice this means that we can have orbits of size $W/k$, where $k$ is a common divisor of $W$ and $R$. In our example with $W=6$ and $R=2,$ we can have orbits of size 6 (when the stabilizer is trivial) or size 3 (when the stabilizer has size 2). We’ve already seen two orbits of size 6, where the stabilizer subgroup is trivial. But for the next starting position, {0,3}, the stabilizer subgroup has two elements: the identity as well as the \(x + 3 \bmod 6\) operation. This means that the resultant orbit will be of size 6/2=3: {0,3}, {1,4}, {2,5}. We can see that this orbit fills each well once rather than twice.
And how well does this approach work in practice? We can compare it to our random baseline from before. Here's a plot comparing the two approaches for $W=24$ and $R=2,3$:
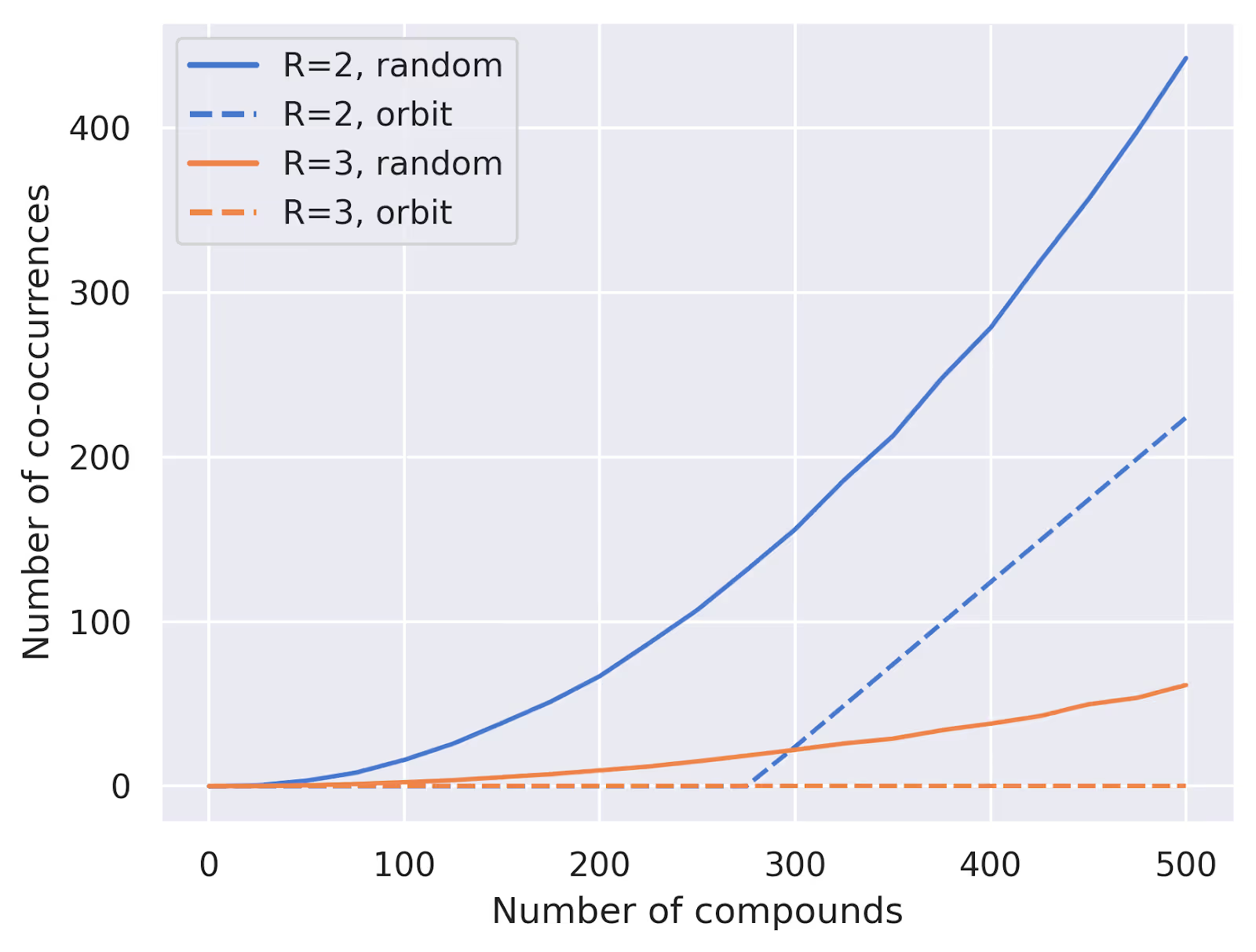
As expected, our systematic approach guarantees no overlaps until we exceed the total number of unique combinations, while the random approach starts to show overlaps much earlier. We can also see that, using this method, we never have a difference between most- and least-filled well greater than $R$:
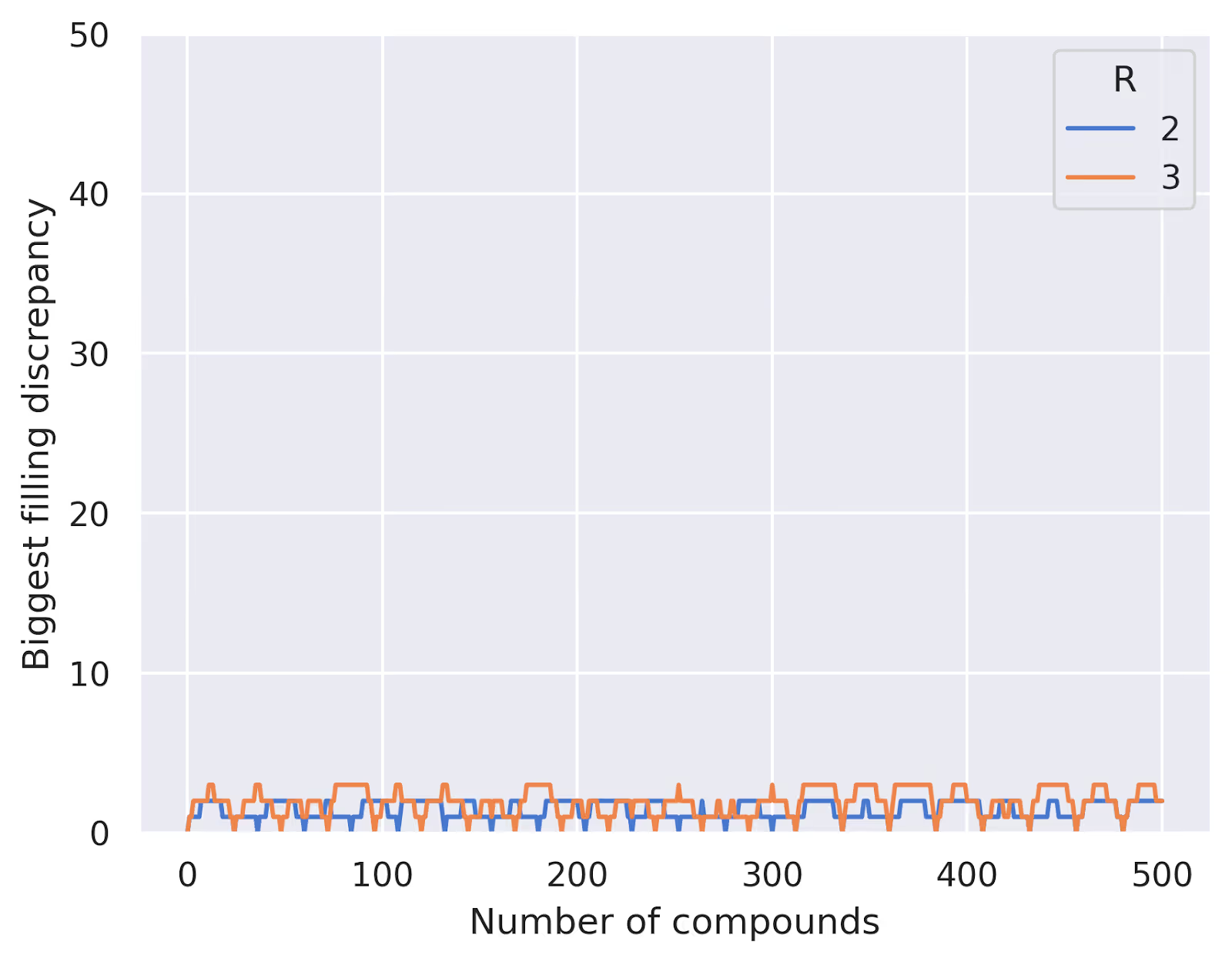
This problem turns out to be closely related to some well-studied and more complex mathematical problems, such as balanced incomplete block designs (BIBDs). And in the real world, very similar ideas arise in contexts like disease testing, error correcting codes, and more. In our case, we're not too concerned with achieving perfect balance between all the wells or conclusively identifying all drug-drug interactions, but this fairly simple approach gives us a practical and efficient way to design our pooling schemes. (We should say, too, that there are lots of other factors that need to be considered when designing a real pooling scheme, but this approach is a good starting point.)
You might think all this math language is a bit of overkill for the problem at hand, and indeed you don't need to understand anything about stabilizers or group actions to implement this solution (though the formal approach is helpful in understanding how to generalize from $R=2$ to larger $R$). But whether we frame the problem with abstract math or not, optimizing our pooling scheme means we can get more data with fewer measurements. And in a field like drug discovery where there are so many difficult trade-offs, it's always nice to find a free win!